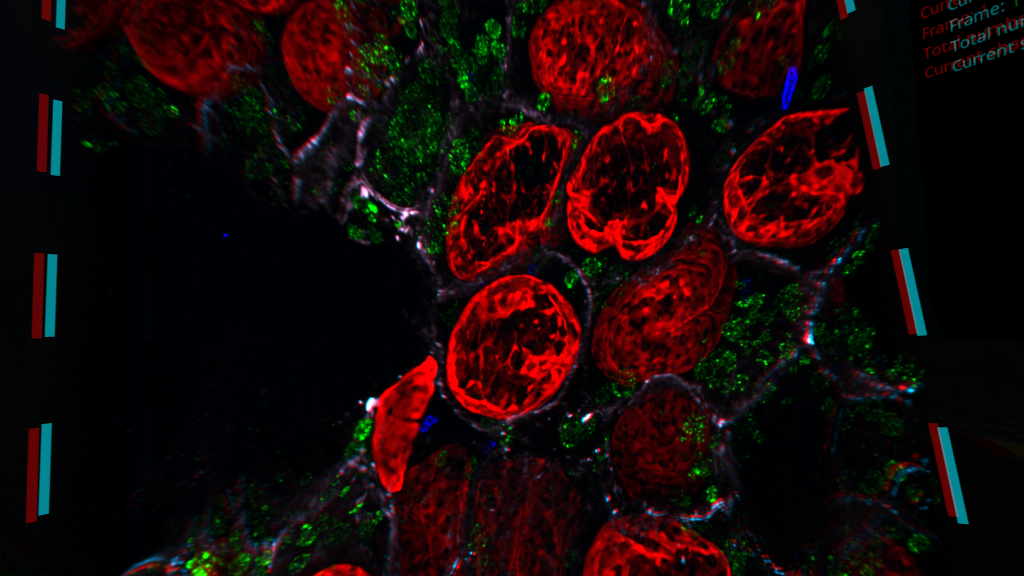
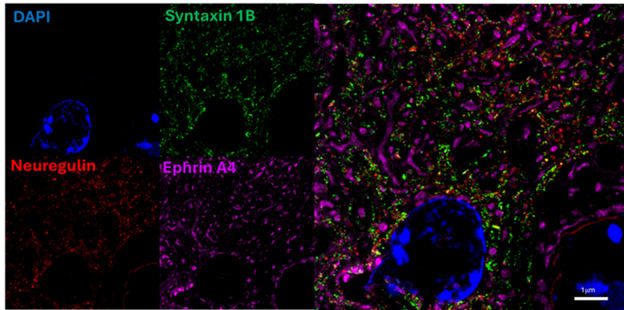
Our core technology Magnify™ has been published in the top-tier peer-reviewed journal Nature Biotechnology!
https://www.nature.com/articles/s41587-022-01546-1
Update: The paper has been downloaded over 15K times in three months and highlighted by Nature Methods and Nature Biomedical Engineering Review, among many news outlets!